Work Packages
WP1 – Infrastructure integration/capacity building: National Cancer Biobank and database
Research Lead: Professor Leonie Young, RCSI
Deputy Lead: Professor Michael Kerin, NUIG

Optimal patient profiling and predictive models of therapeutic response are key to the successful utilisation of targeted therapies. International gap analyses have revealed that translational work in this space has been hampered by a lack of large-scale, standardised and well-annotated breast cancer patient collections, particularly longitudinally sampled patients. As part of this workpackage, we will develop a comprehensive pathophysiological and epidemiological resource to advance the study of tumour progression and ultimately facilitate evidence-based, precision breast cancer medicine. To achieve this, we will consolidate two complementary and pivotal infrastructural components, namely the National Breast Cancer Biobank and the Integrated National Breast Cancer Database, which will be linked to form an integrated National Breast Cancer Biobank and Database. This resource will be invaluable for the population-based, translational and clinical research programmes in WP2-5.
WP2 – Uncovering pathways from population drug exposure to breast cancer outcomes
Research Lead: Professor Kathleen Bennett, RCSI/TCD
Deputy Lead: Dr. Annette Byrne, RCSI

Over 1 in 3 women with breast cancer take aspirin, a statin, or both, prior to their breast cancer diagnosis. Preclinical and epidemiologic data from BREAST-PREDICT investigators and others indicate that these pre-diagnostic drug exposures can have significant effects on breast tumour biology at diagnosis, response to treatment and prognosis for these women. Here, we will utilise the data and bioresources described in WP1 to elucidate the mechanisms by which aspirin and statins influence breast cancer outcome through molecular epidemiologic analyses and pre-clinical studies.
WP3 – Defining the molecular evolution of breast tumors during therapy
Research Lead: Professor Bryan Hennessy, RCSI
Deputy Lead: Professor Des Higgins, UCD
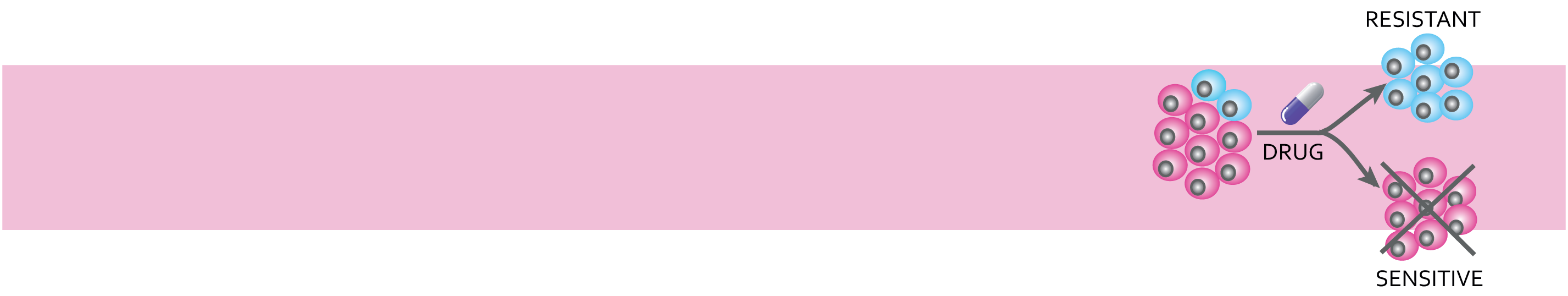

It has long been recognised that molecular differences exist between breast tumours from different patients, even within specific breast cancer subtypes (so called inter-tumour heterogeneity). Moreover, clonal evolution within individual patient cancers leads to intra-tumour heterogeneity, a concept that distinct sub-populations of cancer cells exist within tumours of individual patients, each with distinct genotypes and phenotypes. Although well recognised, clonal evolution and intra-tumour heterogeneity are not well defined in breast cancer subtypes during/after treatment with targeted therapies. In addition, these phenomena are now thought to be intimately involved in tumour progression, the evolution of metastatic disease and treatment resistance. Previous research from BREAST-PREDICT investigators has revealed significant alterations in gene expression between primary, nodal and metastatic tumour tissue in patients who have become resistant to endocrine therapy. This clearly illustrates the plastic nature of breast cancer during therapy and highlights the potential role of prolonged endocrine treatment on the alteration of gene expression. Here, we will use the comprehensive breast bioresources delivered in WP1 to examine the evolution of breast tumours on or after treatment with targeted therapies via pan-omic profiling.
WP4 – Reconstruction of signal transduction networks for the identification of drug targets and rational therapeutic combinations
Research Lead: Professor John Crown, DCU
Deputy Lead: Professor Walter Kolch, UCD

Here, we will leverage the multi-dimensional epidemiologic, pharmacological, genome and proteome data collected in WP1-3 to develop a framework that (1) integrates these diverse datasets; and (2) allows us specifically address molecular mechanisms of drug resistance and how we can overcome them. The strategy is based on a systems biology-driven approach using computational modelling and simulation to analyse highly complex biological data sets and extract the salient mechanistic understanding of drug action and drug resistance. Systems biology approaches will expedite the development of such integrated strategies through predictive modelling of outcomes. As a reflection of this concept, this work package contains global unbiased experimental screening approaches and hypothesis-driven targeted investigations.
WP5 – Pathophysiological validation of signature-based prognostic and predictive biomarkers
Research Lead: Professor Jochen Prehn, RCSI
Deputy Lead: Professor Lorraine O’Driscoll, TCD
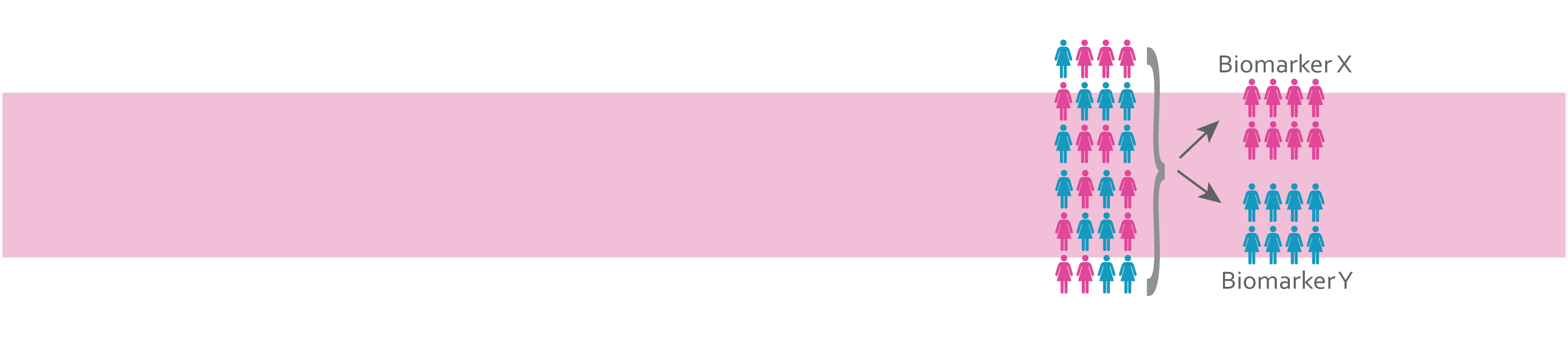

The emergence of multi-marker panels, for example OncotypeDx, now approved by the National Cancer Control Programme for use in Irish breast cancer patients to determine the need for adjuvant chemotherapy in LN- breast cancer patients, and MammaPrint, the first and only breast cancer recurrence test with FDA IVDMIA clearance, highlights the necessity to examine data on a systems level and the power of signature-based diagnostics to tailor therapy. Previous research from BREAST-PREDICT investigators has demonstrated the translational potential of integrated molecular profiling, mathematical modelling and computational biology to identify biomarkers that may guide therapeutic decision-making. In this workpackage, we will clinically verify these signature-based prognostic and predictive biomarker panels (both tissue-based and circulating) using patient samples from a combination of retrospective bioresources (the National Breast Cancer Biobank established in WP1) and ongoing ICORG-led prospective clinical trials.
Our Partners